Online first
Current issue
Archive
Special Issues
About the Journal
Publication Ethics
Anti-Plagiarism system
Instructions for Authors
Instructions for Reviewers
Editorial Board
Editorial Office
Contact
Reviewers
All Reviewers
2025
2024
2023
2022
2021
2020
2019
2018
2017
2016
General Data Protection Regulation (RODO)
RESEARCH PAPER
Detection of Legionella spp. and occurrence of virulence genes: lvh, rtxA and enhC in water samples from artificial water systems
1
Department of Health Biohazards and Parasitology, Institute of Rural Health, Lublin, Poland
Corresponding author
Anna Sawczyn-Domańska
Department of Health Biohazards and Parasitology, Institute of Rural Health, Lublin, Jaczewskiego 2, 20-090, Lublin, Poland
Department of Health Biohazards and Parasitology, Institute of Rural Health, Lublin, Jaczewskiego 2, 20-090, Lublin, Poland
Ann Agric Environ Med. 2021;28(4):617-620
KEYWORDS
TOPICS
ABSTRACT
Introduction and objective:
Legionella spp. are ubiquitous worldwide in natural water sources (rivers, lakes) as well as in man-made water systems (cooling towers, tap water, spas). The most common species causing the disease in humans is Legionella pneumophila. Pathogenicity of Legionella is related to the genes associated with virulence. The study aimed to detect Legionella spp. DNA in man-made water systems, and to examine the presence of selected virulence genes (lvh, rtxA, enhC) in Legionella-positive samples.
Material and methods:
A total of 52 water samples from hot and cold water systems collected from urban and rural areas were investigated for the presence of Legionella by polymerase chain reaction (PCR). The detection of three virulence loci (lvh, rtxA, and enhC) in water samples was investigated using PCR.
Results:
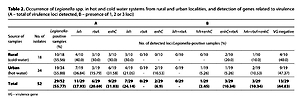
Legionella spp. was detected in over half the tested samples (55.77%), both in water from hot (55.88%) and cold (55.56%) water systems. Sequencing confirmed the presence of Legionella pneumophila in the tested samples. Among all Legionella-positive isolates, at least one virulence gene was detected in the case of 55.17% (16/29) of environmental samples. The lvh locus was most frequently present in all specimens.
Conclusions:
The presence of Legionella pneumophila, the causative agent of Legionnaires’ disease, in hot and cold water systems from rural and urban localities, indicates a risk to public health. The occurrence of virulence genes in tested samples suggests that L. pneumophila has the potential to cause human disease.
Legionella spp. are ubiquitous worldwide in natural water sources (rivers, lakes) as well as in man-made water systems (cooling towers, tap water, spas). The most common species causing the disease in humans is Legionella pneumophila. Pathogenicity of Legionella is related to the genes associated with virulence. The study aimed to detect Legionella spp. DNA in man-made water systems, and to examine the presence of selected virulence genes (lvh, rtxA, enhC) in Legionella-positive samples.
Material and methods:
A total of 52 water samples from hot and cold water systems collected from urban and rural areas were investigated for the presence of Legionella by polymerase chain reaction (PCR). The detection of three virulence loci (lvh, rtxA, and enhC) in water samples was investigated using PCR.
Results:
Legionella spp. was detected in over half the tested samples (55.77%), both in water from hot (55.88%) and cold (55.56%) water systems. Sequencing confirmed the presence of Legionella pneumophila in the tested samples. Among all Legionella-positive isolates, at least one virulence gene was detected in the case of 55.17% (16/29) of environmental samples. The lvh locus was most frequently present in all specimens.
Conclusions:
The presence of Legionella pneumophila, the causative agent of Legionnaires’ disease, in hot and cold water systems from rural and urban localities, indicates a risk to public health. The occurrence of virulence genes in tested samples suggests that L. pneumophila has the potential to cause human disease.
ACKNOWLEDGEMENTS
The study was funded by Institute of Rural Health in Lublin
within a subsidy from the Ministry of Science and Higher
Education in Warsaw, Poland (Project No. 15001).
REFERENCES (32)
1.
Cunha BA, Burillo A, Bouza E. Legionnaires’ disease. Lancet. 2016; 387(10016): 376–385. https://doi.org/10.1016/S0140-....
2.
Phin N, Parry-Ford F, Harrison T, et al. Epidemiology and clinical management of Legionnaires’ disease. Lancet Infect Dis. 2014; 14(10): 1011–1021. https://doi.org/10.1016/S1473-....
3.
The European Legionnaires’ Disease Surveillance Network; The European Centre for Disease Prevention and Control. Surveillance Atlas of Infectious Diseases. Available online: https://atlas.ecdc.europa.eu/p... (access: 2021.10.03).
4.
Miyashita N, Higa F, Aoki Y, et al. Distribution of Legionella species and serogroups in patients with culture-confirmed Legionella pneumonia. Infect Chemother. 2020; 26(5): 411–417. https://doi.org/10.1016/j.jiac....
5.
Zhan XY, Zhu QY. Molecular typing of Legionella pneumophila isolates from environmental water samples and clinical samples using a five-gene sequence typing and standard Sequence-Based Typing. PLoS One. 2018; 13(2): e0190986. https://doi.org/10.1371/journa....
6.
Potočnjak M, MagdaleniĆ Z, Dijan M, et al. Environmental factors affecting the survival of soil dwelling Legionella longbeachae in water. Ann Agric Environ Med. 2016; 23(3): 452–455. https://doi.org/10.5604/123219....
7.
van Heijnsbergen E, Schalk JA, Euser SM, et al. Confirmed and potential sources of Legionella reviewed. Environ Sci Technol. 2015; 49(8): 4797–4815. https://doi.org/10.1021/acs.es....
8.
Prussin AJ, Schwake DO, Marr LC. Ten questions concerning the aerosolization and transmission of Legionella in the built environment. Build Environ. 2017; 123: 684–695. https://doi.org/10.1016/j.buil....
9.
Khweek AA, Amer A. Replication of Legionella pneumophila in human cells: Why are we susceptible?. Front Microbiol. 2010; 1: 133. https://doi.org/10.3389/fmicb.....
10.
Samrakandi MM, Cirillo SL, Ridenour DA, et al. Genetic and phenotypic differences between Legionella pneumophila strains. J Clin Microbiol. 2002; 40(4): 1352–1362. https://doi.org/10.1128/JCM.40....
11.
Bandyopadhyay P, Liu S, Gabbai CB, et al. Environmental mimics and the Lvh type IVA secretion system contribute to virulence-related phenotypes of Legionella pneumophila. Infect Immun. 2007; 75(2): 723–735. https://doi.org/10.1128/IAI.00....
12.
Cirillo SLG, Yan L, Littman M, et al. Role of the Legionella pneumophila rtxA gene in amoebae. Microbiology. 2002; 148(Pt 6): 1667–1677. https://doi.org/10.1099/002212....
13.
Liu M, Conover GM, Isberg RR. Legionella pneumophila EnhC is required for efficient replication in tumour necrosis factor alpha-stimulated macrophages. Cell Microbiol. 2008; 10(9): 1906–1923. https://doi.org/10.1111/j.1462....
14.
Wójcik-Fatla A, Stojek NM, Dutkiewicz J. Efficacy of the detection of Legionella in hot and cold water samples by culture and PCR. I. Standardization of methods. Ann Agric Environ Med. 2012; 19(2): 289–293.
15.
Sikora A, Wójtowicz-Bobin M, Kozioł-Montewka M, et al. Prevalence of Legionella pneumophila in water distribution systems in hospitals and public buildings of the Lublin region of eastern Poland. Ann Agric Environ Med. 2015; 22(2): 195–201. https://doi.org/10.5604/123219....
16.
Wojtyła-Buciora P, Chrzanowska E, Marcinkowski JT. Presence of Legionella sp. in hot water systems of health care institutions and public buildings. Hygeia Public Health 2013; 48(3): 327–332.
17.
Napoli C, Fasano F, Iatta R, et al. Legionella spp. and legionellosis in southeastern Italy: disease epidemiology and environmental surveillance in community and health care facilities. BMC Public Health. 2010; 10: 660. https://doi.org/10.1186/1471-2....
18.
Żak J, Orlińska K, Koperny M, et al. Legionella sp. in water systems in public teaching and education facilities in Małopolskie voivodeship in 2016. Przegl Epidemiol. 2019; 73(2): 227–237. https://doi.org/10.32394/pe.73....
19.
Pierre D, Baron JL, Ma X, et al. Water quality as a predictor of Legionella positivity of building water systems. Pathogens. 2019; 8(4): 295. https://10.3390/pathogens80402....
20.
Farnham A, Alleyne L, Cimini D, et al. Legionnaires’ disease incidence and risk factors, New York, New York, USA, 2002–2011. Emerg Infect Dis. 2014; 20(11): 1795–1802. https://doi.org/10.3201/eid201....
21.
Lampl BMJ, Lang M, Wodnick S. Can mandatory monitoring in rental apartments effectively prevent legionellosis? A retrospective analysis of data from Regensburg with a review of the literature. GMS Hyg Infect Control. 2020; 15: Doc14. https://doi.org/10.3205/dgkh00....
22.
Ricci ML, Rota MC, Caporali MG, et al. A Legionnaires’ disease cluster in a private building in Italy. Int J Environ Res Public Health. 2021; 18(13): 6922. https://doi.org/10.3390/ijerph....
23.
Hayes-Phillips D, Bentham R, Ross K, et al. Factors influencing Legionella contamination of domestic household showers. Pathogens. 2019; 8(1): 27. https://doi.org/10.3390/pathog....
24.
Huang B, Heron BA, Gray BR, et al. A predominant and virulent Legionella pneumophila serogroup 1 strain detected in isolates from patients and water in Queensland, Australia, by an amplified fragment length polymorphism protocol and virulence gene-based PCR assays. J Clin Microbiol. 2004; 42: 4164–4168. https://doi.org/10.1128/JCM.42....
25.
Sreenath K, Chaudhry R, Vinayaraj EV, et al. Distribution of virulence genes and sequence-based types among Legionella pneumophila isolated from the water systems of a tertiary care hospital in India. Front Public Health. 2020; 8: 596463. https://doi.org/10.3389/fpubh.....
26.
Katsiaflaka A, Pournaras S, Kristo I, et al. Epidemiological investigation of Legionella pneumophila serogroup 2 to 14 isolates from water samples by amplified fragment length polymorphism and sequence-based typing and detection of virulence traits. Appl Environ Microbiol. 2016; 82: 6102–6108. https://doi.org/10.1128/AEM.01....
27.
Hwang IY, Park EH, Lee YC. Distribution of virulence genes in environmental and clinical isolates of Legionella pneumophila in Busan, South Korea. Southeast Asian J Trop Med Public Health. 2017; 48(1): 83–90.
28.
Toplitsch D, Platzer S, Zehner R, et al. Comparison of updated methods for Legionella detection in environmental water samples. Int J Environ Res Public Health. 2021; 18(10): 5436. https://doi.org/10.3390/ijerph....
29.
Alleron L, Khemiri A, Koubar M, et al. VBNC Legionella pneumophila cells are still able to produce virulence proteins. Water Res. 2013; 47(17): 6606–6617. https://doi.org/10.1016/j.watr....
30.
Dietersdorfer E, Kirschner A, Schrammel B, et al. Starved viable but non-culturable (VBNC) Legionella strains can infect and replicate in amoebae and human macrophages. Water Res. 2018; 141: 428–438. https://doi.org/10.1016/j.watr....
31.
Collins S, Jorgensen F, Willis C, et al. Real-time PCR to supplement gold-standard culture-based detection of Legionella in environmental samples. J Appl Microbiol. 2015; 119(4): 1158–1169. https://doi.org/10.1111/jam.12....
32.
National Institute of Public Health-National Institute of Hygiene, 2021. Reports on cases of infectious diseases and poisonings in Poland. http://www.pzh.gov.pl (access: 2021.10.03).
Share
RELATED ARTICLE
We process personal data collected when visiting the website. The function of obtaining information about users and their behavior is carried out by voluntarily entered information in forms and saving cookies in end devices. Data, including cookies, are used to provide services, improve the user experience and to analyze the traffic in accordance with the Privacy policy. Data are also collected and processed by Google Analytics tool (more).
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.